Code
load("../data/train.dtf.Rdata")
load("../data/test.dtf.Rdata")January 24, 2024
Création de jeux de données corrects
#train.names <- sample(rownames(X), 2*nrow(X)/3, replace = FALSE)
#test.names <- rownames(X)[-which(rownames(X) %in% train.names)]
#res.sample <- PCA(X, scale.unit = TRUE, ind.sup = match(test.names, rownames(X)))
#fviz_pca_biplot(res.sample, geom='point')
#train.set <- X[train.names,]
#test.set <- X[test.names,]
#train.dtf <- cbind(train.set, Y[train.names,])
#test.dtf <- cbind(test.set, Y[test.names,])
#save(train.dtf, file = "../data/train.dtf.Rdata")
#save(test.dtf, file = "../data/test.dtf.Rdata")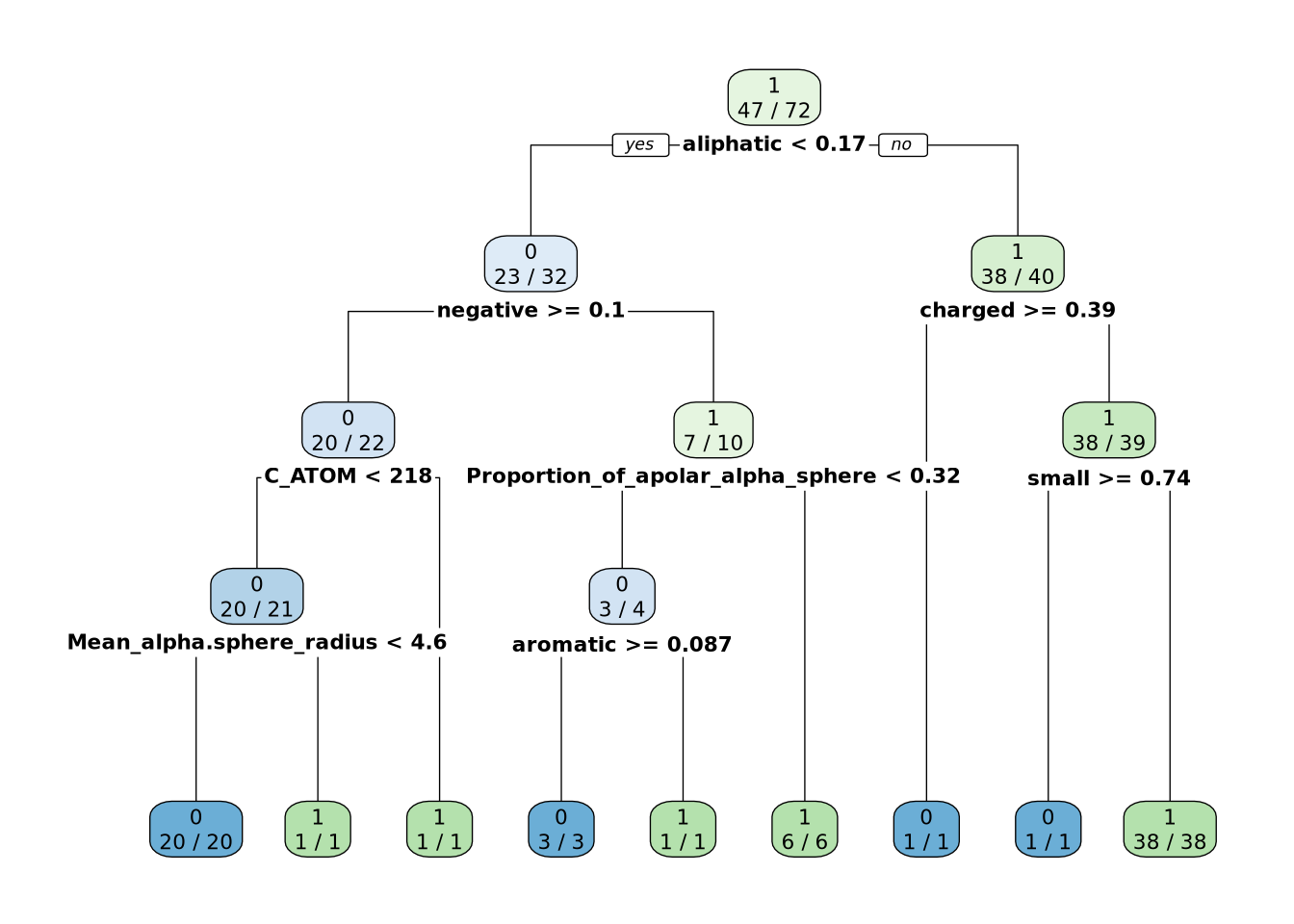
Loading required package: ggplot2Loading required package: lattice# Train set
fit_predict_train <- predict(Fit, newdata = train_drugg)
conf_train_fit <- confusionMatrix(as.factor(round(fit_predict_train[, 2])),
as.factor(train_drugg$drugg))
# Test set
fit_predict_test <- predict(Fit, newdata = test_drugg)
conf_test_fit <- confusionMatrix(as.factor(round(fit_predict_test[, 2])),
as.factor(test_drugg$drugg))
# Plot
plotcp(Fit)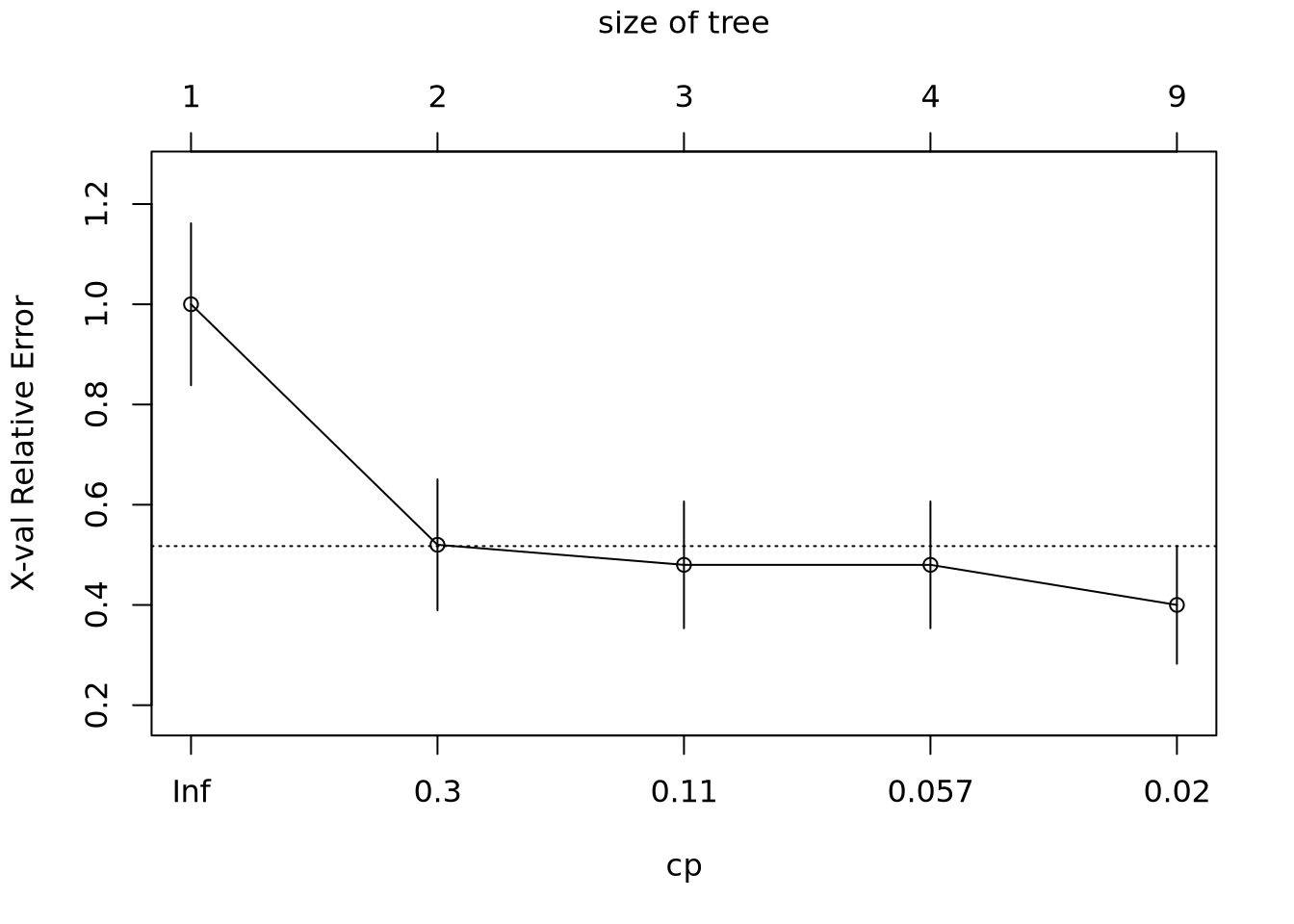
# Train set
ness_predict_train <- predict(Ness, newdata = train_drugg)
conf_train_ness <- confusionMatrix(as.factor(round(ness_predict_train[, 2])),
as.factor(train_drugg$drugg))
# Test set
ness_predict_test <- predict(Ness, newdata = test_drugg)
conf_test_ness <- confusionMatrix(as.factor(round(ness_predict_test[, 2])),
as.factor(test_drugg$drugg))Accuracy
1 Accuracy
0.7027027 Accuracy
0.9305556 Accuracy
0.7027027 Accuracy
train_fit 1.0000000
test_fit 0.7027027
train_ness 0.9305556
test_ness 0.7027027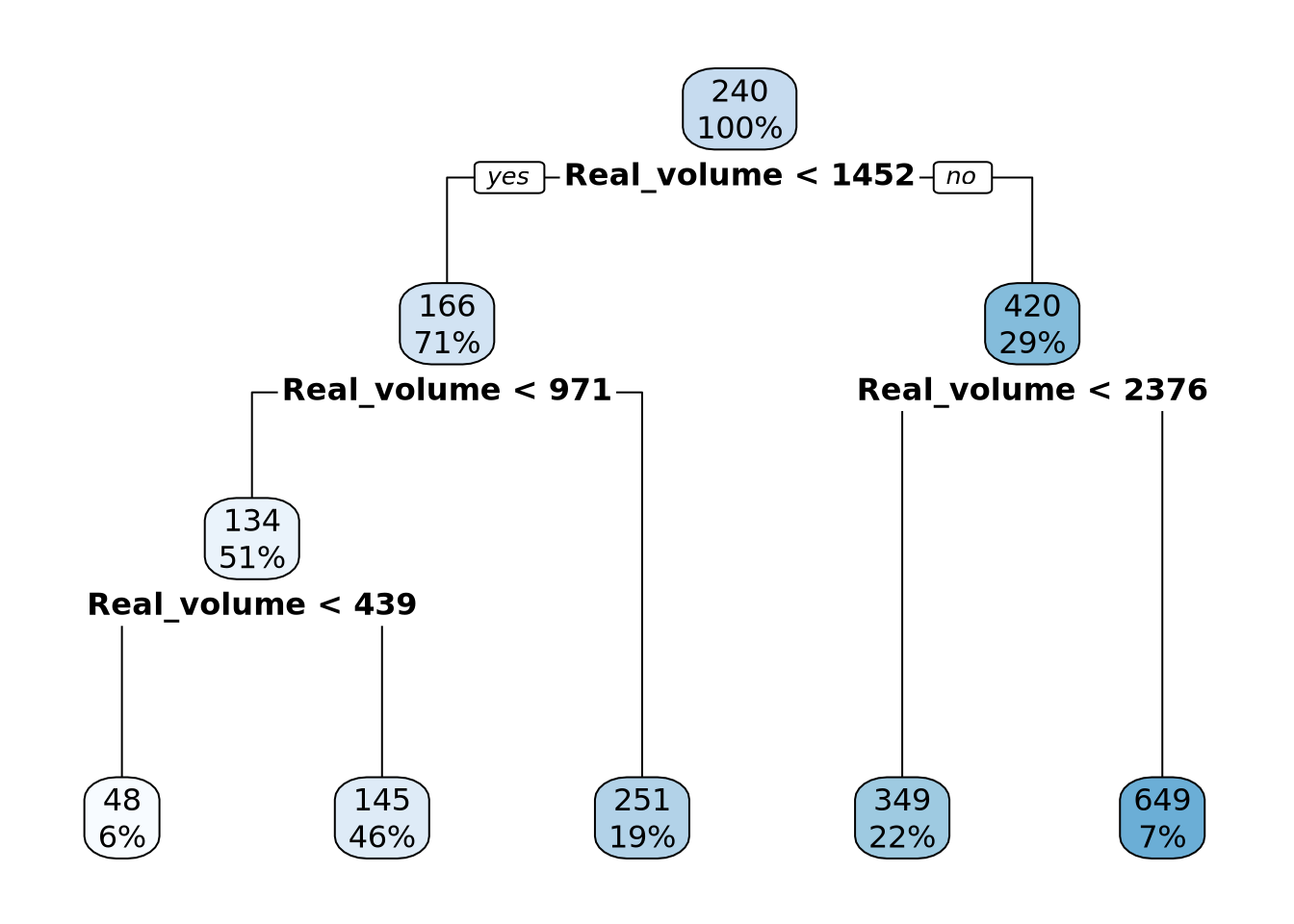
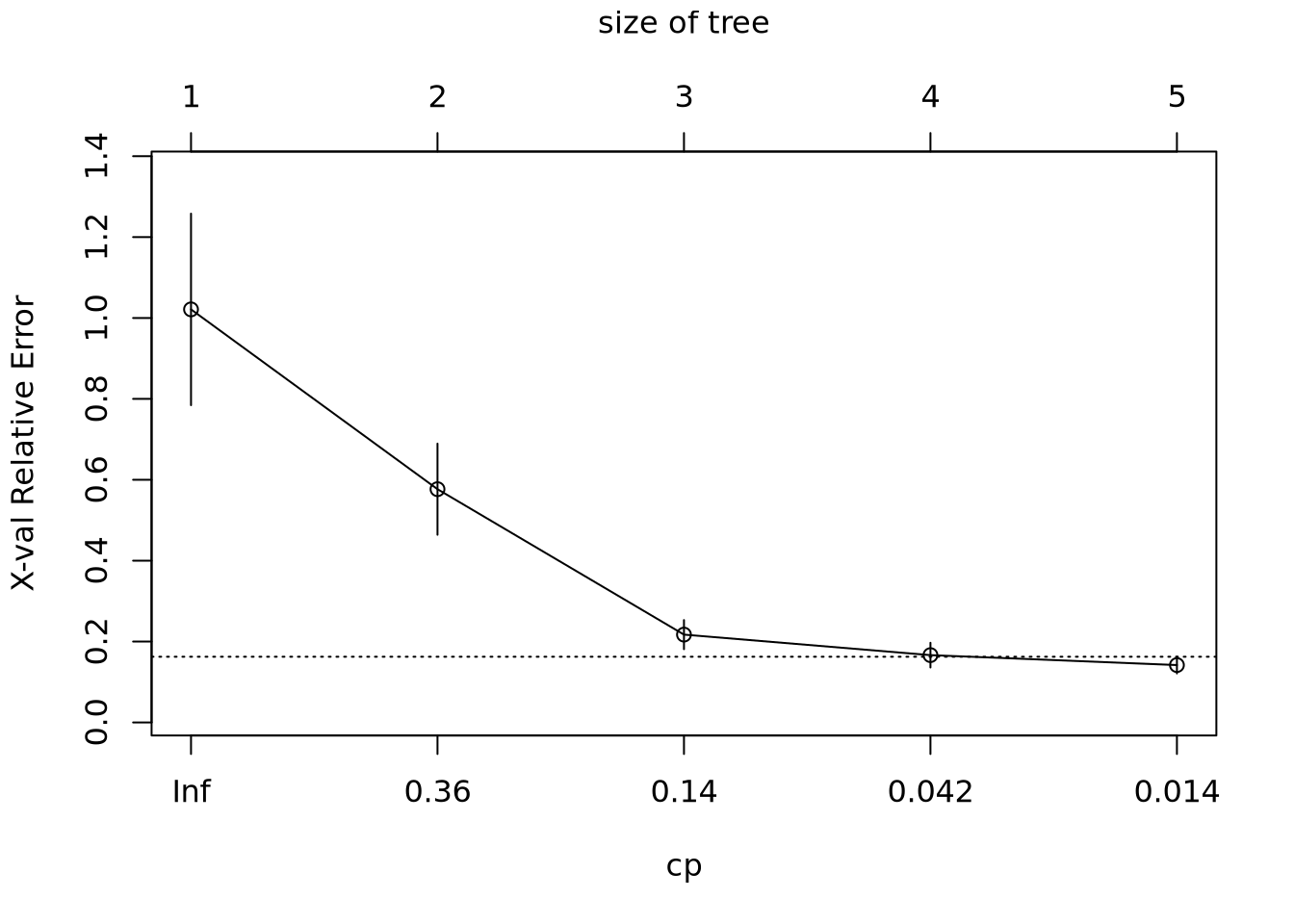
# Create a dataframe
results_df <- data.frame(
RMSE = c(41.3386484, 50.6410931, 46.5379404, 50.6410931),
Rsquared = c(0.9229773, 0.9299857, 0.9023842, 0.9299857),
MAE = c(33.3168518, 39.2805625, 37.7836200, 39.2805625),
row.names = c("train_original", "test_original", "train_pruned", "test_pruned")
)@online{bari garnier2024,
author = {Bari Garnier, Martin},
title = {Partitioning {Trees}},
date = {2024-01-24},
url = {https://MartinBaGar.github.io/Master_ISDD_fiches//mda/pages/tp4.html},
langid = {en}
}